Band Structure of 1D Lattice
Plotting electronic band structure is one of many exciting things in condensed matter physics. Acquiring the skill to calculate and plot the band structure will open the way to study many things. In this tutorial, we will learn how to calculate and plot a band structure using Python and Julia. Before tackling models with complex lattice structure, we take a simple 1-dimensional lattice as our first baby step.
A normal Hamiltonian for a single particle is written like this,
\[H(x) = -\frac{\hbar^2}{2m}\frac{d^2}{dx^2} + V(x).\]We can think of a lattice as a system with periodic potential, so
\[V(x + a) = V(x),\]with some period \(a\).
Lattice Vectors
Our system consists of equally spaced points, so the lattice vecors are just integer \(n\) times the spacing \(a\):
\[R_n = na.\]What about the reciprocal lattice vectors \(G_n\)? Recall that in general,
\[\vec{G} \cdot \vec{R} = 2\pi m\]where \(m\) is an integer. In our case:
\[G R = Ga = 2\pi \Rightarrow G = \frac{2\pi}{a}\]which means
\[G_n = \frac{2\pi}{a}n.\]Applying Bloch’s Theorem
Next, let’s apply Bloch’s theorem to our system. According to Bloch’s theorem, the wavefunction of a periodic system can be expressed as
\[\psi_k(x) = e^{i k x} u_k(x)\]where \(u_k(x)\) is periodic,
\[u_k(x + a) = u_k(x)\]Now, we calculate \(\frac{d^2}{dx^2} \psi_k(x)\) using this:
\[\begin{split} \frac{d^2}{dx^2} \psi_k(x) &= \frac{d}{dx} \left( \frac{d}{dx} e^{i k x} u_k(x) \right)\\ &=\frac{d}{dx} \left( e^{ikx} \frac{d}{dx} u_k(x) + ik e^{ikx} u_k(x) \right)\\ &=ik e^{ikx} \frac{d}{dx} u_k(x) + e^{ikx} \frac{d^2}{dx^2} u_k(x) + ike^{ikx} \frac{d}{dx}u_k(x) + i^2 k^2 e^{ikx} u_k(x)\\ &=e^{ikx} \left(\frac{d^2}{dx^2} + 2 ik \frac{d}{dx} + i^2k^2 \right) u_k(x)\\ &=e^{ikx} \left(\frac{d}{dx} + ik \right)^2 u_k(x) \end{split}\]So then, the eigenvalue equation of our system can be written as
\[\begin{split} H \psi_k(x) &= E \psi_k(x) \\ \left[ -\frac{\hbar^2}{2m} \left(\frac{d}{dx} + ik \right)^2 + V \right] u_k(x) &= E u_k(x) \end{split}\]If you want, you can also consider that \(p = -i\hbar \frac{d}{dx}\) and then rewrite the equation as
\[\left[ \frac{\hbar^2}{2m} \left(p + \hbar k \right)^2 + V \right] u_k(x) = E u_k(x)\]For now, let’s just set \(\hbar = 1\) so we have a simple looking equation
\[\boxed{\left[ \frac{1}{2m} \left(p + k \right)^2 + V \right] u_k(x) = E u_k(x) }\]Expanding the Hamiltonian
Now, we will expand our Hamiltonian in some basis. Firstly, let us set the potential as the following:
\[V(x) = v \cos(G_1 x)\]where \(v\) is a constant which determines the strength of the potential. So, our Hamiltonian becomes
\[H = \frac{1}{2m} \left(p + k \right)^2 + v \cos(G_1 x)\]We choose plane waves as our basis:
\[u_k(x) = \frac{1}{\sqrt{a}}e^{i G_{\mu} x}\]and then expand our Hamiltonian in this basis, so the matrix element is:
\[\begin{split} H_{\mu\nu} &= \frac{1}{a} \int_0^a e^{-iG_{\mu} x} \left[ \frac{1}{2m} \left(-i\frac{d}{dx} + k \right)^2 + v \cos(G_1 x) \right] e^{i G_{\nu} x} dx \\ &= \frac{1}{a} \int_0^a e^{-i \frac{2\pi}{a}{\mu} x} \left[ \frac{1}{2m} \left(-\frac{d^2}{dx^2} -2i k \frac{d}{dx} + k^2 \right) + v \cos(G_1 x) \right] e^{i \frac{2\pi}{a}{\nu} x} dx \end{split}\]We have two derivatives, let’s calculate them one by one.
- First term:
- Second term:
Let’s also look at the potential term:
\[\begin{split} v \cos(G_1x) e^{i\frac{2\pi}{a}(\nu)x} &= \frac{v}{2}\left(e^{iG_1 x} + e^{-iG_1x}\right) e^{i\frac{2\pi}{a}(\nu)x} \\ &= \frac{v}{2} \left( e^{i \frac{2\pi}{a}x } e^{i\frac{2\pi}{a}(\nu)x} + e^{-i \frac{2\pi}{a}x } e^{i\frac{2\pi}{a}(\nu)x} \right) \\ &= \frac{v}{2} \left( e^{i\frac{2\pi}{a}(\nu + 1)x} + e^{i\frac{2\pi}{a}(\nu - 1)x} \right) \end{split}\]Now, we have:
\[\begin{split} H_{\mu\nu} (k) &= \frac{1}{a} \int e^{-i\frac{2\pi}{a}\mu x} \frac{1}{2m} \left[ \left( \frac{2\pi}{a} \nu \right)^2 + \frac{4\pi}{a}\nu k + k^2 \right] e^{i\frac{2\pi}{a}\nu x} dx \\ &\quad \, + \frac{1}{a} \int e^{-i\frac{2\pi}{a}\mu x} \frac{v}{2} \left( e^{i\frac{2\pi}{a}(\nu + 1)x} + e^{i\frac{2\pi}{a}(\nu - 1)x} \right) dx \\ &= \frac{1}{2m} \left( \frac{2\pi}{a} \nu + k \right)^2 \frac{1}{a} \int e^{i\frac{2\pi}{a}(\nu-\mu) x} dx \\ &\quad \, + \frac{v}{2} \left( \frac{1}{a} \int e^{i\frac{2\pi}{a}(\nu + 1 - \mu)x} dx + \frac{1}{a} \int e^{i\frac{2\pi}{a}(\nu - 1 - \mu)x} dx \right) \end{split}\]We can then use the identity:
\[\frac{1}{a} \int_{0}^{a} e^{i(G_{\nu}-G_{\mu})x} dx = \delta_{\nu,\mu}\]where \(\delta_{i,j}\) is a Kronecker delta. Here, we used \(G_n = \frac{2\pi}{a}n\) so that
\[G_{\nu}-G_{\mu} = \frac{2\pi}{a}(\nu - \mu).\]Using this identity, the matrix element becomes
\[\boxed{H_{\mu\nu} (k) = \frac{1}{2m} \left( \frac{2\pi}{a} \nu + k \right)^2 \delta_{\nu, \mu} + \frac{v}{2} \left(\delta_{\nu,\mu - 1} + \delta_{\nu,\mu + 1} \right)}\]This is the equation that is useful for our numerical computation. What a long discussion. So, what did we do so far? In summary, after applying Bloch’s theorem, we rewrote the Hamiltonian at a fixed crystal momentum \(k\) and expanded the periodic part of the wavefunction \(u_k(x)\) in a plane-wave basis. What we have then is a matrix eigenvalue problem for each \(k\).
In practice, we truncate the number of plane waves by restricting \(\mu\) to a finite range so we keep a finite number of Fourier components of \(u_k(x)\). Of course, increasing the number of basis states improve the accuracy of the calculation.
Let’s Feed the Computer
Now is time to put our equation into action. I will show how to calculate and plot the band structure in Python and Julia. You can use whichever language you like. It’s useful to use Jupyter notebook in this stage so we can see the result of any calculation directly. For Python, we will need Numpy package so let’s call it first:
import numpy as np
For Julia, we will use LinearAlgebra package:
using LinearAlgebra
Let’s also define our constants. To make it simple, we will just set \(m\) and \(a\) to 1. The following is valid for both Python and Julia:
m = 1.0
a = 1.0
v = 0.1
Next, it is useful to write a helper function for Kronecker delta. In Python:
def kronDelta(i, j):
return i == j
In Julia, we can write
function kronDelta(i, j)
return i == j
end
Some people might think this is unnecessary, that’s fine though, but I like Kronecker deltas.
It’s time to write our Hamiltonian. I will try to keep the same symbol as we have in our equation.
def hamiltonian(m, a, v, mu, nu, k):
return (0.5 / m) * ( 2 * np.pi * nu / a + k ) ** 2 * kronDelta(nu, mu) + (v / 2) * (kronDelta(nu, mu - 1) + kronDelta(nu, mu + 1))
In Julia:
function hamiltonian(m, a, v, mu, nu, k)
return (0.5 / m) * ( 2 * pi * nu / a + k )^2 * kronDelta(nu, mu) + (v / 2) * (kronDelta(nu, mu - 1) + kronDelta(nu, mu + 1))
end
Next, let’s define our \(k\) points which are inside the first Brillouin zone. Because we have a 1D system, the first Brillouin zone is just a line spanning from \(-\pi/a\) to \(\pi/a\) in the \(k\) space. So, we now make a list of number in that range. In Python, we can use Numpy’s linspace:
kpoints = np.linspace(-np.pi/a, np.pi/a, 100)
In Julia, there is a similar function called LinRange,
kpoints = LinRange(-pi/a, pi/a, 100)
Note that the above functions give us a list of 100 numbers, ranging from \(-\pi/a\) to \(\pi/a\) with equal interval.
We will also make a list of basis index. They are just integers and we can make them in both Python and Julia using list comprehension.
In Python:
basis_idx = [i for i in range(-2, 3)]
And in Julia we write:
basis_idx = [i for i in -2:2]
Okay, time to compute the eigenvalue. Here is what we will do: For one \(k\)-point, we expand the Hamiltonian using the plane wave basis then get the eigenvalue of the Hamiltonian for that particular \(k\)-point. We then move to the next \(k\)-point and do the same thing again until the last \(k\)-point in our list.
To begin, we will make an empty list called ergs as a container for the eigenvalues. Then, we loop through the list of the \(k\)-points. In each iteration, we first make a square matrix filled with zero. This will be our Hamiltonian matrix. We then loop through the list of the basis index and construct the Hamiltonian. After we get the full Hamiltonian matrix. We take the eigenvalues and store them in ergs.
This is how we write it in Python:
N = len(basis_idx)
ergs = []
for k in kpoints:
H = np.zeros((N, N))
for mu in range(N):
for nu in range(N):
H[mu, nu] = hamiltonian(m, a, v, basis_idx[mu], basis_idx[nu], k)
ergs.append(np.linalg.eigvalsh(H))
ergs = np.array(ergs)
Note that for our convenience, we convert ergs which was initially a list to a Numpy array. We also take an advantage by the fact that our Hamiltonian is a real and symmetric matrix so we use Numpy’s linalg.eigvalsh function instead of the normal linalg.eigvals.
In Julia, we have this:
N = length(basis_idx)
ergs = Vector{Vector{Float64}}()
for k in kpoints
H = zeros(N, N)
for mu in 1:N
for nu in 1:N
H[mu, nu] = hamiltonian(m, a, v, basis_idx[mu], basis_idx[nu], k)
end
end
push!(ergs, eigvals(Symmetric(H)))
end
See also that indexing is different in Python and Julia, so just be careful. You can also see that we have tagged the H matrix as symmetric by imposing the function Symmetric so that Julia can do operation on it more efficiently.
Great! Now we just need to plot the eigenvalues. Let’s do it first in Python.
We call Matplotlib package for plotting:
import matplotlib.pyplot as plt
Because ergs already contains eigenvalues for each \(k\)-point, the plotting is straightforward:
fig = plt.figure(figsize=(8, 6))
plt.plot(kpoints, ergs)
plt.xlabel("k")
plt.ylabel("Energy")
plt.show(fig)
This is what we get in Python:
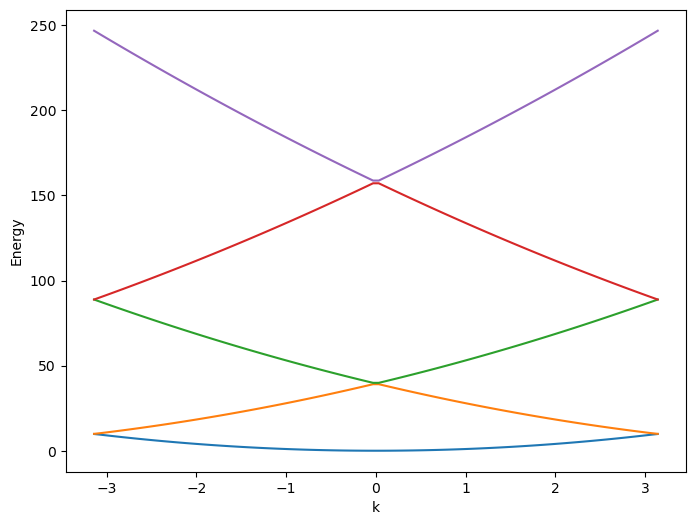
Let’s do it also in Julia. Our container ergs is basically a list containing eigenvalues for each \(k\)-point. It will be convenient to plot the band structure by converting ergs to a matrix. This can be done using stack() function. For plotting, I will show it using CairoMakie.jl.
ergs_matrix = Float64.(stack(ergs)) # Convert ergs to a matrix of (num_of_bands, num_of_kpoints)
num_bands = size(ergs_matrix, 1) # Number of bands for each k
fig = Figure(size=(800,600))
ax = Axis(fig[1,1], xlabel="k-path", ylabel="Energy", xlabelsize=24, ylabelsize=24)
for j in 1:num_bands
lines!(ax, kpoints, ergs_matrix[j, :])
end
fig
This is what we get:
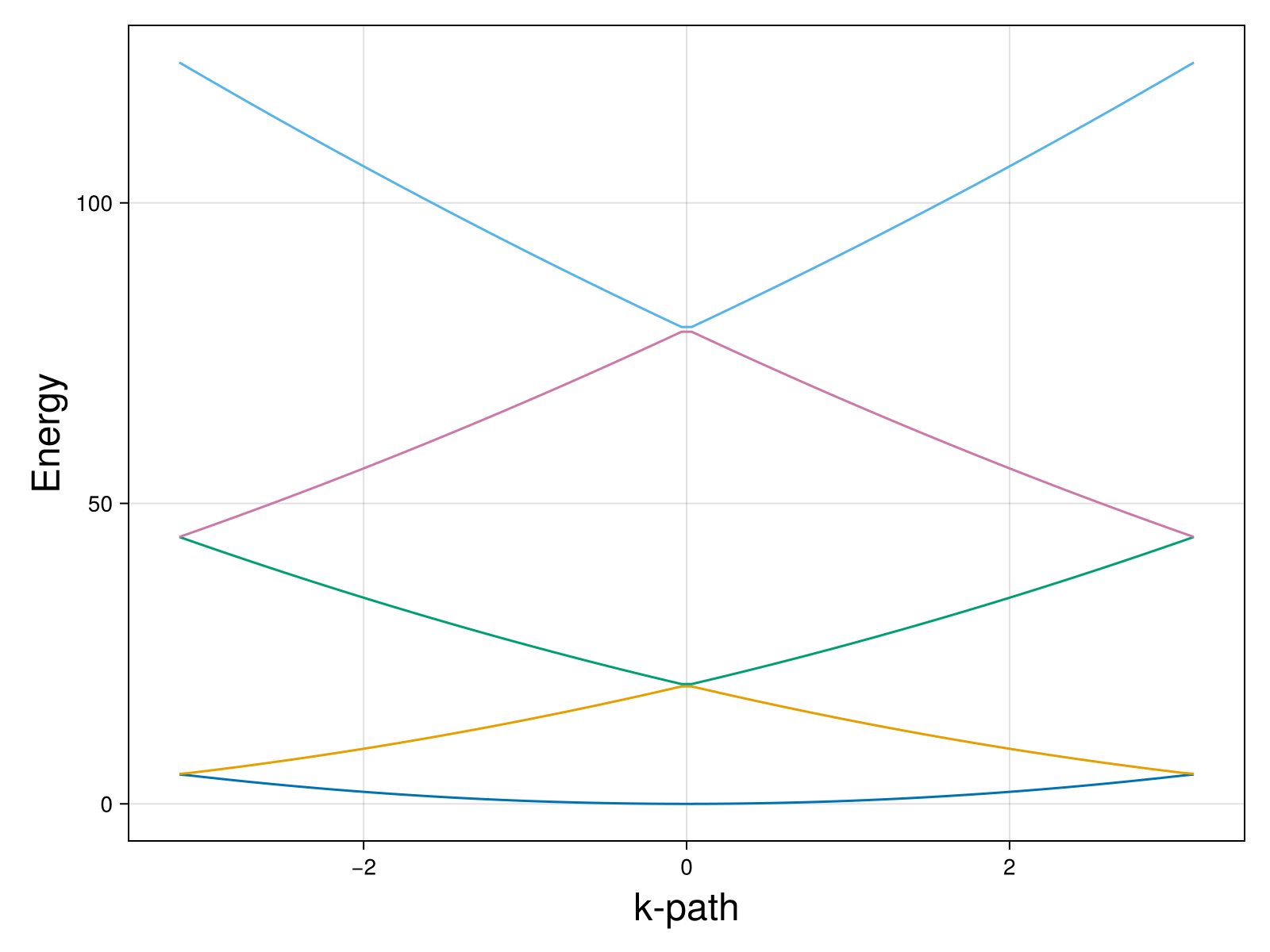
And we are done. You can play with the constant to see the change in the band structure. So far, we set v = 0.1 which makes this practically an empty lattice. If you set v = 10, for example, you will see a gap opening in the band structure:
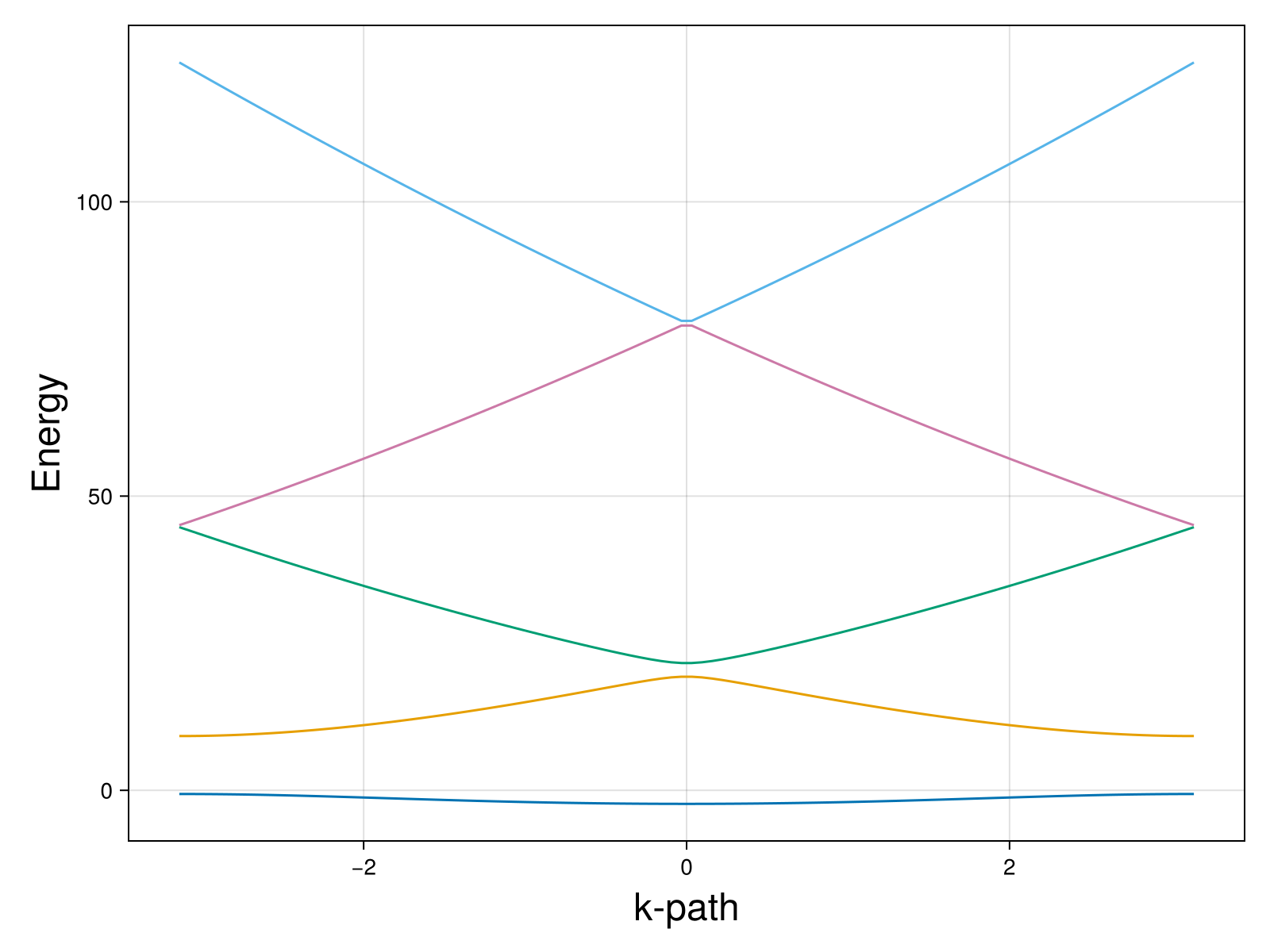
Phew, it’s a long post. I hope you enjoy it. You can also view the code here: